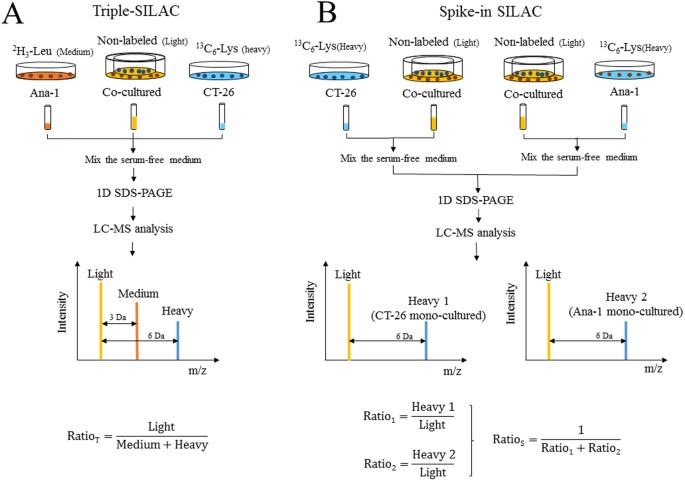
Use of stable isotope labeling by amino acids in cell culture as a spike-in standard in quantitative proteomics | Nature Protocols
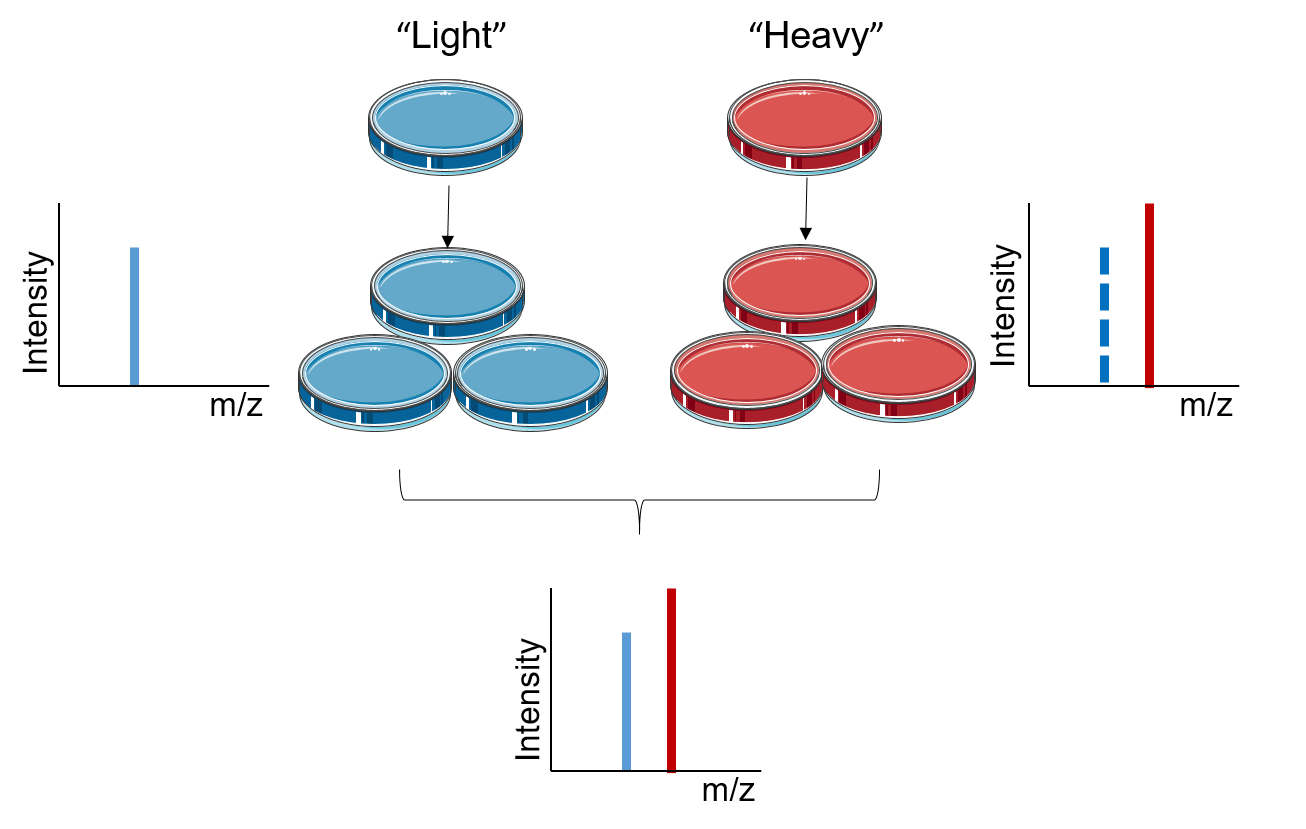
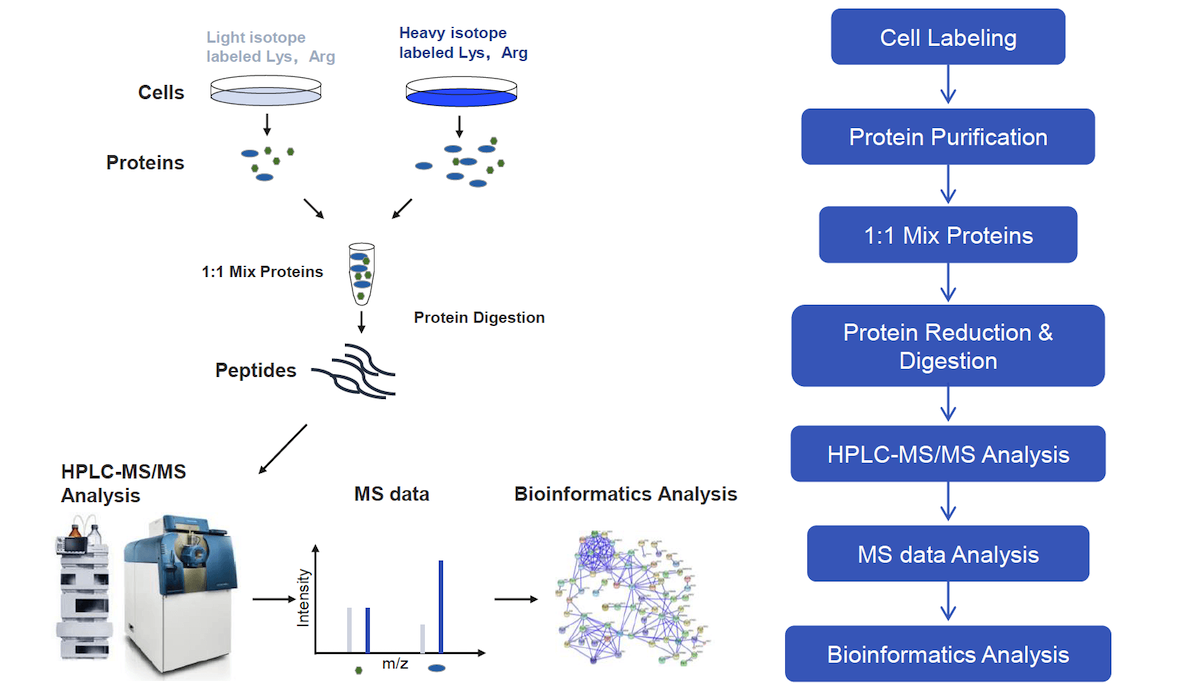
Stable Isotope Labeling Using Amino Acids in Cell Culture (SILAC): Principles, Workflow, and Applications – Creative Proteomics Blog

Determination of an Angiotensin II-regulated Proteome in Primary Human Kidney Cells by Stable Isotope Labeling of Amino Acids in Cell Culture ( SILAC) - Journal of Biological Chemistry
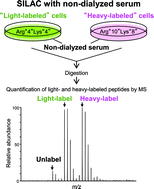
Quantitative proteome and phosphoproteome analyses of cultured cells based on SILAC labeling without requirement of serum dialysis - Molecular BioSystems (RSC Publishing)

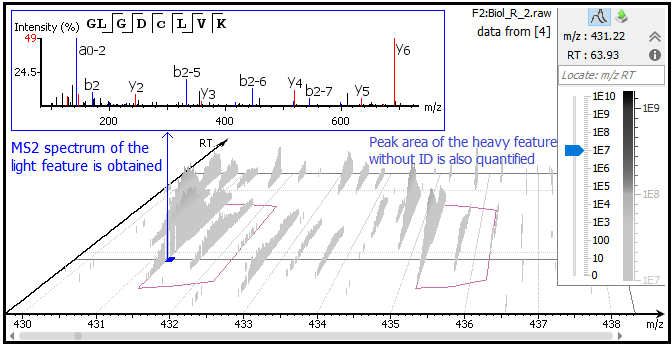
Use of stable isotope labeling by amino acids in cell culture (SILAC) for phosphotyrosine protein identification and quantitation. - Abstract - Europe PMC

Stable Isotope Labeling by Amino Acids in Cell Culture (SILAC) for Quantitative Proteomics | SpringerLink

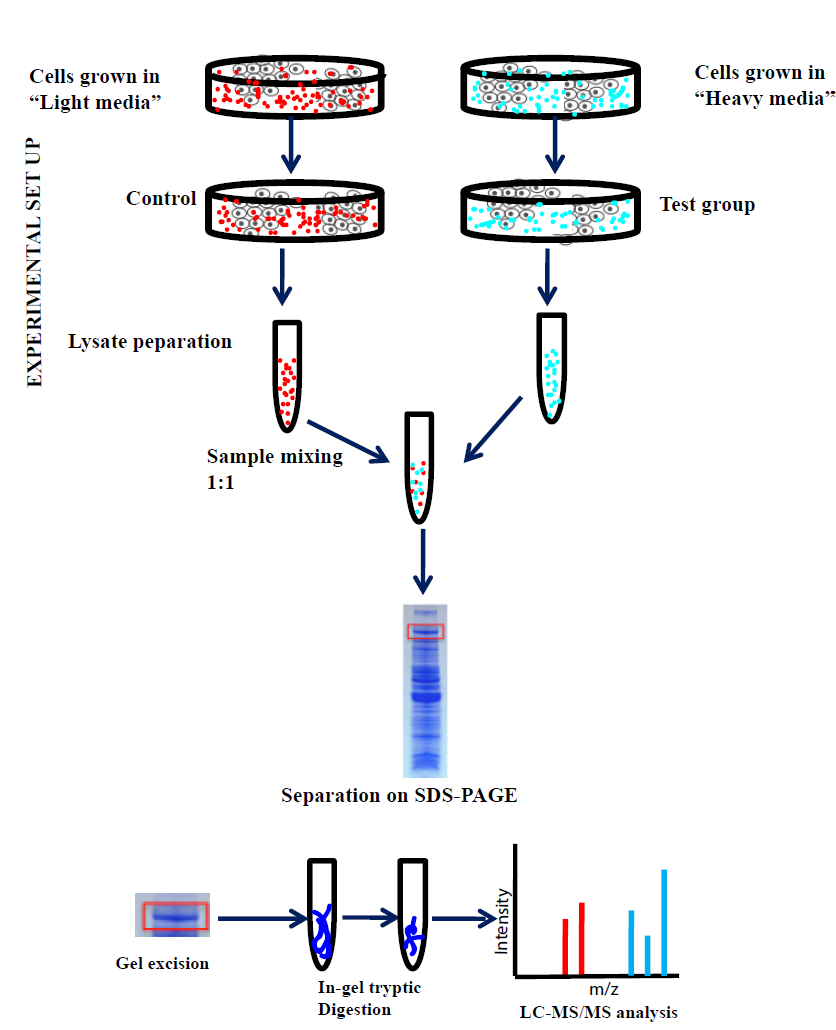
Stable Isotope Labeling by Amino Acids in Cell Culture, SILAC, as a Simple and Accurate Approach to Expression Proteomics* - Molecular & Cellular Proteomics

JCI Insight - Deep phenotyping of human induced pluripotent stem cell–derived atrial and ventricular cardiomyocytes

IJMS | Free Full-Text | SILAC-Based Quantification of TGFBR2-Regulated Protein Expression in Extracellular Vesicles of Microsatellite Unstable Colorectal Cancers | HTML

Quantitative proteomics using SILAC: Principles, applications, and developments - Chen - 2015 - PROTEOMICS - Wiley Online Library
PLOS ONE: Semi-quantitative proteomics of mammalian cells upon short-term exposure to non-ionizing electromagnetic fields